As the COVID-19 pandemic scattered and isolated people, researchers across Virginia Tech connected for a data-driven collaboration seeking improved drugs to fight the disease and potentially many other illnesses.
A multidisciplinary collaboration spanning several colleges at Virginia Tech resulted in a newly published study, “Data Driven Computational Design and Experimental Validation of Drugs for Accelerated Mitigation of Pandemic-like Scenarios,” in the Journal of Physical Chemistry Letters.
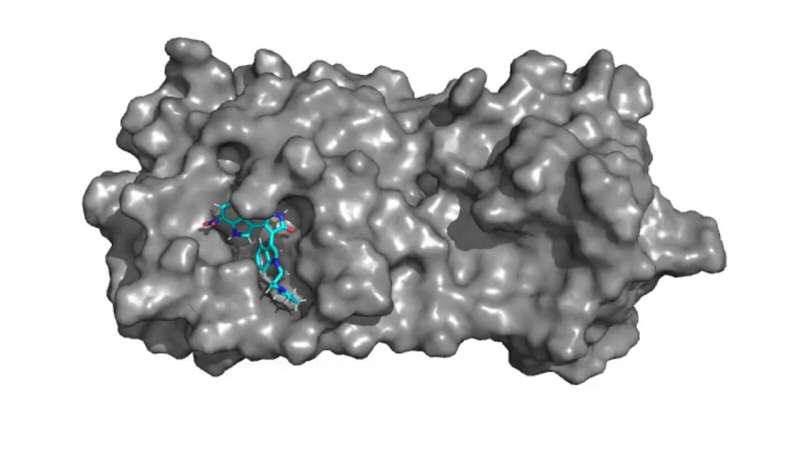
The study focuses on using computer algorithms to generate adaptations to molecules in compounds for existing and potential medications that can improve those molecules’ ability to bind to the main protease, a protein-based enzyme that breaks down complex proteins, in SARS-CoV-2, the virus that causes COVID-19.
This process allows exponentially more molecular adaptations to be considered than traditional trial-and-error methods of testing drugs one by one could allow. Candidate molecule adaptations can be identified among myriad possibilities, then narrowed to a few or one that can be created in a laboratory and tested for effectiveness.
“We present a novel transferable data-driven framework that can be used to accelerate the design of new small molecules and materials, with desired properties, by changing the combination of building blocks as well as decorating them with functional groups,” said Sanket A. Deshmukh, associate professor of chemical engineering in the College of Engineering. A “functional group” is a cluster of atoms that generally retains its characteristic properties, regardless of the other atoms in the molecule.
“Interestingly, the newly designed functionalized drug not only had a better half maximal effective concentration value than its parent drug, but also several of the proposed and used antivirals, including remdesivir,” Deshmukh said, referring to a measure of compound potency.
Moving through all the phases of the study would not have been possible without extensive cross-departmental collaboration.

Four Virginia Tech faculty members—Deshmukh; Anne M. Brown, associate professor with University Libraries and the Department of Biochemistry in the College of Agriculture and Life Sciences; Andrew Lowell, assistant professor in the Department of Chemistry in the College of Science; and James Weger-Lucarelli, assistant professor in the Department of Biomedical Sciences and Pathobiology at the Virginia-Maryland College of Veterinary Medicine—are among 13 co-authors of the published study.
The Deshmukh group’s expertise in developing transferable computational models and frameworks for accelerated design of drug-like small molecules and materials, and Brown’s extensive computational expertise in protein structure-function relationships, meshed seamlessly as a baseline for the study.
“Sanket’s group had a molecular repurposing framework, and I have experience with exploiting protein targets,” said Brown. “Combined with Andrew, who does the synthesis, which is to make the compound, and then James doing the testing and the viral assays, we formed a fantastic collaboration.”
But the faculty members stress that it was the graduate students and postdoctoral students in the laboratories who made the study possible. Nine of them are co-authors: Samrendra K. Singh, chemical engineering; Kelsie King, genetics, bioinformatics, and computational biology; Cole Gannett, chemistry; Christina Chuong, biomedical and veterinary sciences; Soumil Y. Joshi, chemical engineering; Charles Plate, chemical engineering; Parisa Farzeen, chemical engineering; Emily M. Webb, entomology; and Lakshmi Kumar Kunche, chemical engineering.
The professors said the students communicated well with one another without any prompting from their mentors. “I think one of the great things to see is the students really talking with one another and collaborating with one another as well without us having to say ‘Do this,'” Deshmukh said.
Finally, the functionalized molecules were tested against live SARS-CoV-2 in a veterinary college laboratory by Weger-Lucarelli and his team.
“Initial virtual screening of the existing database identified a parent compound that was expected to inhibit the protease of SARS-CoV-2,” Weger-Lucarelli said. “Then the data-driven framework altered the structure of that molecule to enhance that activity. We compared those side by side to show that this new compound that was expected to be more potent against SARS-CoV-2 than the parent compound was, in fact, more potent against SARS-CoV-2.”

The process to develop and test a functionalized molecule against COVID-19 has many potential applications even beyond mitigation of COVID-19. Studies are ongoing among the team to employ the same type of research to find functionalized molecules that may be able to treat hepatitis E, dengue fever and chikungunya, the latter two being mosquito-borne illnesses.
“Another direction we’re going in is that we’re targeting proteases and enzymes from other viruses and trying to design other new molecules,” Lowell said.
The algorithm process also has potential in non-biological uses, Sankit said. The “approach is very versatile and is being applied to functionalize and design other materials such as metal organic frameworks (MOFs), glycomaterials, polymers, etc.,” the paper states.
The assembled interdisciplinary team is planning to continue its collaborations.
“None of us could do this work without the other people in this collaboration,” Weger-Lucarelli said.
“This is a great example of the synergy between going from computational prediction to chemical synthesis to testing in viruses,” Brown said, “and how we at Virginia Tech are really emphasizing that interplay between these three areas and taking that to the next level to develop strong collaborative teams.”
More information: Data Driven Computational Design and Experimental Validation of Drugs for Accelerated Mitigation of Pandemic-like Scenarios, The Journal of Physical Chemistry Letters (2023).
Journal information: Journal of Physical Chemistry Letters
Provided by Virginia Tech

